2021年1月20日,北京大兴区通报了2例本土新冠肺炎病例样本的病毒基因测序结果,结果表明2例样本均属于L基因型欧洲家系分支(B.1.1.7),与英国发现的新冠病毒突变株高度同源。至此,国内上海、广东和北京等地均已通报检出B.1.1.7突变株感染者,引起了广泛关注。
新冠病毒的突变点都有哪些?突变会导致怎样的影响?日常的核酸检测是否还有效?今天,笔者就带大家详细了解一下新冠病毒的这些突变“兄弟们”。
什么是病毒突变株?
病毒在进行复制时,其遗传物质会发生变化,这些变化被称为“突变”。而具有一个或多个新突变的病毒被称为原始病毒的“变体”(即突变株)。新冠病毒作为一种分子稳定性相对较差的RNA病毒,发生突变是再正常不过的一件事情,目前全球已经发现了数百种新冠病毒的变体,其中大部分对新冠病毒的属性影响很小甚至无影响,我们真正需要关注的是那些会影响病毒属性的突变株。
新冠病毒的“有效”突变株主要从两方面来影响病毒的属性:
1)增强新冠病毒与宿主蛋白ACE2受体的亲和力,从而增强病毒的感染能力;
2)帮助新冠病毒逃逸宿主免疫系统的攻击,导致人类二次感染概率升高甚至降低疫苗有效性。
目前,尚无证据表明已知的突变点会增强新冠病毒的致病性(人感染后病情加重)。
五种重要的突变株
世界卫生组织(WHO)2020年12月31日正式通报了自新冠病毒疫情爆发以来主要的4种病毒突变株,分别为D614G突变株、“Cluster 5”突变株、B.1.1.7突变株和B.1.351突变株,加上近日“突起”的P.1突变株,共有5种引人注目的新冠突变株。
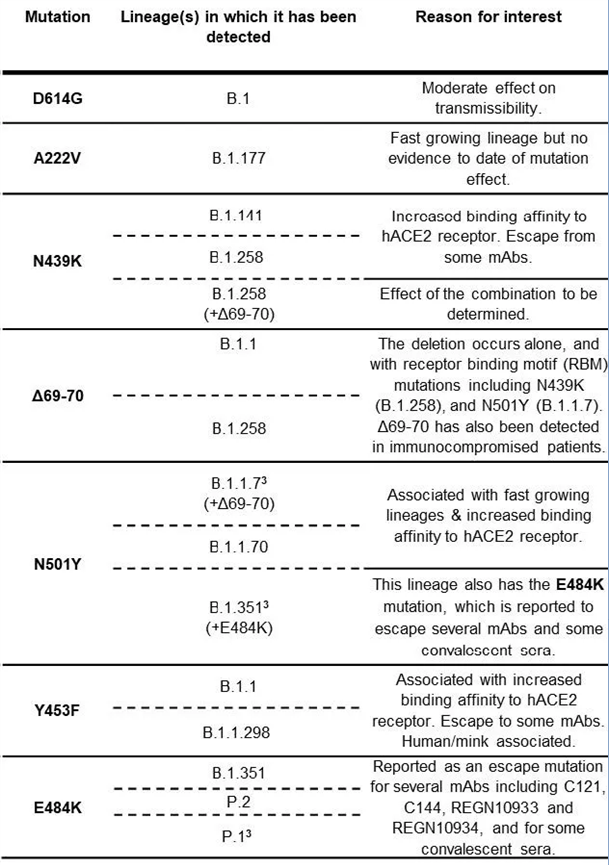
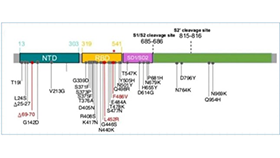
图1 不同突变株需要重点关注的突变位点
●D614G突变株
2020年1月底发现的D614G突变株,也称B.1突变株,其突变位点主要为S蛋白上的D614G。至2020年6月,D614G突变株已成为全球范围内流行的主要毒株,其占比高达64.6%。D614G突变株的复制和传播更快,感染能力更强,传染性更强。
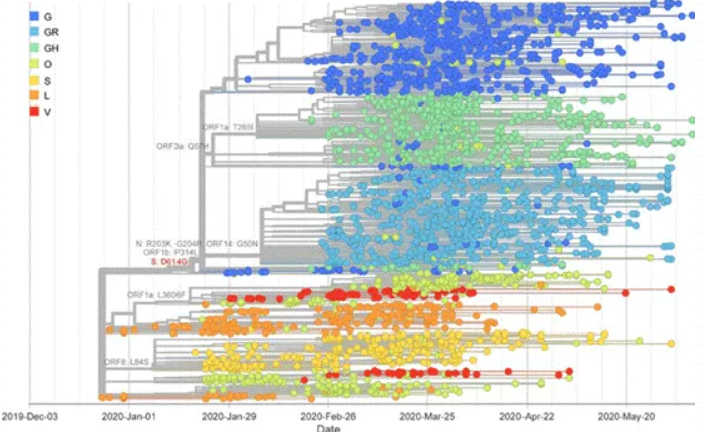
图2 D614突变株的流行趋势
●“Cluster 5”突变株
“Cluster 5”突变株是人与中间宿主互相传播的典型代表,其可在人与水貂中传播。“Cluster 5”突变株于2020年8-9月在丹麦发现,至少含有2个关键突变:Y453F突变和69-70del突变。世界卫生组织称,“Cluster 5”突变株可能造成人体中和抗体的活性适度减弱,导致人体在自然感染或接种疫苗后,产生的免疫保护范围和持续时间缩短,但其在世界范围内流行程度较低。
●B.1.1.7突变株
2020年12月14日英国向世卫组织通报出现新的“B.1.1.7突变株”(又称VUI-202012/01),该毒株的3个位于S基因上的关键突变具有潜在的生物学功能,分别为N501Y突变、69-70del突变和p681H突变。初步的研究结果表明,B.1.1.7突变株的传染性可能比普通毒株高出56%,目前已传播至约60个国家和地区。
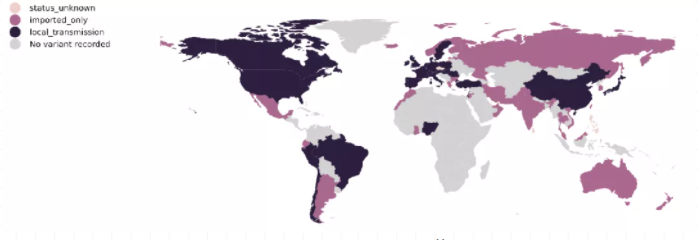
图3 B.1.1.7突变株的全球流行情况
●B.1.351突变株
B.1.351突变株自2020年8月首次发现于南非并迅速传播,于11月初成为南非的主要流行株。B.1.351突变株的3个位于S基因上的关键突变具有潜在的生物学功能,分别为N501Y突变、E484K突变和K417N突变。最新的研究结果表明,B.1.351突变株的传播能力更强,对其他毒株诱导的中和抗体的敏感性显著下降,导致人类二次感染新冠的风险增高。
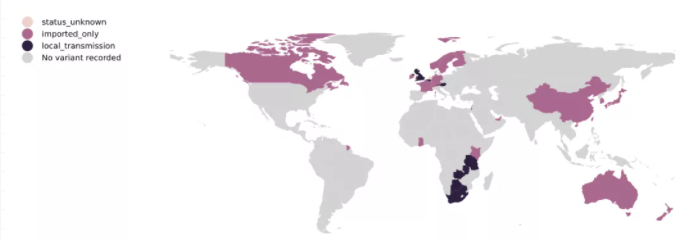
图4 B.1.351突变株的全球流行情况
●P.1突变株
P.1突变株属于B.1.1.28分支,于2021年1月1日在圣保罗被发现,随后传至日本。P.1突变株的3个位于S基因上的关键突变具有潜在的生物学功能,分别为N501Y突变、E484K突变和K417T突变。K417是中和抗体(neutralizing antibodies,nAbs)的重要靶点,有研究表明,该突变可降低病毒与人体内中和抗体的亲和力,从而有助于免疫逃逸。
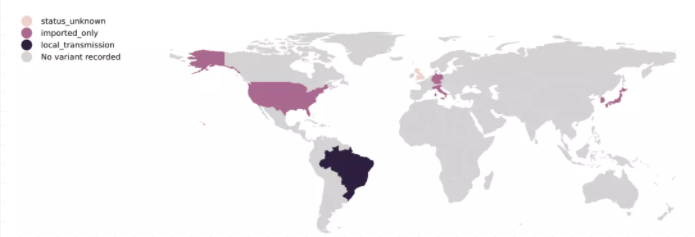
图5 P.1突变株的全球流行情况
莫慌!核酸检测依然靠谱
新型冠状病毒肺炎疫情爆发以来,新冠病毒核酸检测已经成为群众日常生活中司空见惯的场景,甚至在2021年的春节,一张核酸检测阴性报告成为了大家回乡的“车票”。当下,越来越多新冠病毒突变株的出现引起了人们对于新冠核酸检测有效性的担忧。莫慌,笔者这就给大家排解焦虑~
依据国家新冠诊疗方案的要求,国内主流的新冠检测试剂靶标多为ORF1ab基因和N基因,而目前发现的新冠突变株的“有效”突变点基本集中在S蛋白上,对于检测的有效性不会造成显著影响。此外,伯杰医疗目前亦已完成对现有试剂引物探针与突变株的核对,结果证实:伯杰医疗已经上市的新型冠状病毒2019-nCoV核酸检测试剂盒(荧光PCR法),针对最新突变株均不会出现脱靶和漏检,仍可保证检测试剂的准确性和灵敏性。
如何快速鉴别新冠突变株?
截至目前,全球已知的新冠病毒变体高达数百种,其中引人关注的B.1.1.7突变株、B.1.351突变株和P.1突变株存在的免疫逃逸现象已经引起广泛关注和讨论,隐患依然存在。因而,世卫组织及国际专家依然在密切监测新冠病毒的变化,以便能及时发现重大突变,国内也仍需投入巨大的精力监测新冠突变株的流行情况。看来,对于这些突变株,不该省的工序还是不能省~
当下,全球对新冠突变株(尤其是未知突变株)的监测,主要依赖基因测序技术以及全球科研人员对测序结果数据的共享。但基因测序的时间往往较长,在情况紧急时,及时响应成了一个大问题。因而,对于已知有特异性突变点的突变株,我们也可以通过荧光PCR技术对突变株进行快速鉴别分型和溯源监测,避免贻误疫情防控。

参考文献
[1]Zhang L, Jackson C B, Mou H, et al. The D614G mutation in the SARS-CoV-2 spike protein reduces S1 shedding and increases infectivity[J]. bioRxiv, 2020.
[2]Daniloski Z, Guo X, Sanjana N, et al. The D614G mutation in SARS-CoV-2 Spike increases transduction of multiple human cell types[J]. bioRxiv, 2020.
[3] Huang AL, et al. The D614G mutation of SARS-CoV-2 spike protein enhances viral infectivity and decreases neutralization sensitivity to individual convalescent sera. bioRxiv, 2020.
[4]Fonager J., et al., Working paper on SARS-CoV-2 spike mutations arising in Danish mink, their spread to humans and neutralization data. https://files.ssi.dk/Mink-cluster-5-short-report_AFO2. 2020.
[5]van Dorp, L., et al., Recurrent mutations in SARS-CoV-2 genomes isolated from mink point to rapid host-adaptation. bioRxiv, 2020: p. 2020.11.16.384743.
[6]Starr, T.N., et al., Deep Mutational Scanning of SARS-CoV-2 Receptor Binding Domain Reveals
Constraints on Folding and ACE2 Binding. Cell, 2020. 182(5): p. 1295-1310.e20.
[7]Andrew Rambaut, et al. Preliminary genomic characterisation of an emergent SARS-CoV-2 lineage in the UK defined by a novel set of spike mutations. Genomics Consortium UK (CoG-UK), December 19, 2020. Hoffmann et al. A Multibasic Cleavage Site in the Spike Protein of SARS-CoV-2 Is Essential for Infection of Human Lung Cells, 2020, Molecular Cell 78, 779–784, May 21, 2020.
[8]Gu et al. Adaptation of SARS-CoV-2 in BALB/c mice for testing vaccine efficacy, Science 369, 1603–1607 (2020).
[9]Starr et al. Deep Mutational Scanning of SARS-CoV-2 Receptor Binding Domain Reveals Constraints on Folding and ACE2 Binding , 2020, Cell 182, 1295–1310, September 3, 2020.
[10]Kevin R. McCarthy et al. Natural deletions in the SARS-CoV-2 spike glycoprotein drive antibody escape. bioRxiv, December 19, 2020.
[11]Kemp SA, et al. Neutralising antibodies drive Spike mediated SARS-CoV-2 evasion. medRxiv, December 19, 2020.
[12]Thomas P. Peacock, et al. The furin cleavage site of SARS-CoV-2 spike protein is a key determinant for transmission due to enhanced replication in airway cells. bioRxiv, September 30, 2020.
[13]Yunkai Zhu, et al. The S1/S2 boundary of SARS-CoV-2 spike protein modulates cellentry pathways and transmission. bioRxiv, August 25, 2020.
[14]Houriiyah Tegally, et al.Emergence and rapid spread of a new severe acute respiratory syndrome-related coronavirus 2 (SARS-CoV-2) lineage with multiple spike mutations in South Africa. medRxiv, December 21, 2020.
[15]Baum, A., et al., Antibody cocktail to SARS-CoV-2 spike protein prevents rapid mutational escape seen with individual antibodies. Science, 2020: p. eabd0831.
[16]Greaney, A.J., et al., Comprehensive mapping of mutations to the SARS-CoV-2 receptor-binding domain that affect recognition by polyclonal human serum antibodies. bioRxiv, 2021: p.
2020.12.31.425021.
[17]Xie, X., et al., Neutralization of N501Y mutant SARS-CoV-2 by BNT162b2 vaccine-elicited sera. bioRxiv, 2021: p. 2021.01.07.425740.
[18]Rambaut, A., et al., A dynamic nomenclature proposal for SARS-CoV-2 lineages to assist genomic epidemiology. Nature Microbiology, 2020. 5(11): p. 1403-1407.
[19]Faria, N.R., et al. Genomic characterisation of an emergent SARS-CoV-2 lineage in Manaus:
preliminary findings. 2021; Available from: https://virological.org/t/genomic-characterisation-of-anemergent-sars-cov-2-lineage-in-manaus-preliminary-findings/586.
[20]Naveca, F., et al. Phylogenetic relationship of SARS-CoV-2 sequences from Amazonas with emerging Brazilian variants harboring mutations E484K and N501Y in the Spike protein. 2021; Available from: https://virological.org/t/phylogenetic-relationship-of-sars-cov-2-sequences-from-amazonas-withemerging-brazilian-variants-harboring-mutations-e484k-and-n501y-in-the-spike-protein/585.
[21]Voloch, C.M., et al., Genomic characterization of a novel SARS-CoV-2 lineage from Rio de Janeiro, Brazil. medRxiv, 2020: p. 2020.12.23.20248598.上海伯杰医疗科技有限公司是一家致力于感染性病原体分子诊断试剂研发和应用,深耕于多重荧光PCR诊断试剂和痕量病毒二代测序试剂及相关服务的国家高新技术企业。公司围绕感染性病原体这一主线,从诊断试剂、诊断仪器、测序服务和医检所服务等多个面提供全套解决方案。公司秉承“勇于创新,质量为先”的方针,为医疗机构、疾控公卫、高校科研等合作伙伴提供优质产品与服务。

全国客服电话:400-860-3688














.jpg)